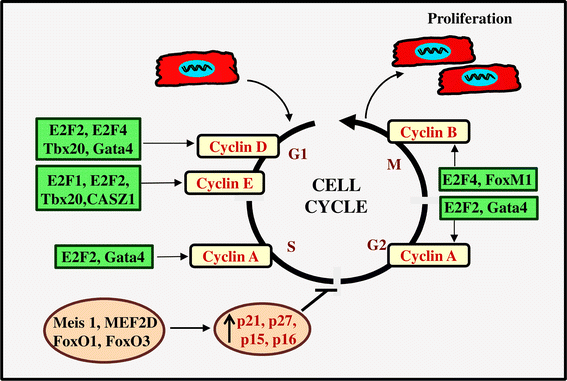
E2F transcription factors regulate expression of a panel of cellular genes that control cellular DNA synthesis
DNA synthesis
DNA synthesis is the natural or artificial creation of deoxyribonucleic acid (DNA) molecules. The term DNA synthesis can refer to DNA replication - DNA biosynthesis (in vivo DNA amplification), polymerase chain reaction - enzymatic DNA synthesis (in vitro DNA amplification) or gene synthesis - physically creating artificial gene sequences.
Full Answer
What is the function of E2F1 in eukaryotes?
E2F is a group of genes that encodes a family of transcription factors (TF) in higher eukaryotes. Three of them are activators: E2F1, 2 and E2F3a. Six others act as suppressors: E2F3b, E2F4-8. All of them are involved in the cell cycle regulation and synthesis of DNA in mammalian cells.
What is the role of E2F and DP in cell cycle?
The E2F, DP, and Rb families play key roles in regulation of the mammalian cell cycle. E2F and DP family members comprise transcription factors that regulate transcription of genes encoding proteins necessary for progression through S phase of the cell cycle, for example, DHFR, Orcl, Myb, Myc, cdc2, and others (56, 57).
What is the function of E2F3a?
E2F is a group of genes that encodes a family of transcription factors (TF) in higher eukaryotes. Three of them are activators: E2F1, 2 and E2F3a.
How does E2F affect transcriptional activity?
The net effect of E2F on the transcriptional activity of a particular gene may be the result of as yet poorly understood protein-protein interactions of E2F with other components of the transcriptional machinery, as well as it may reflect the readout of the different ways of regulating E2F activity, itself.

What does the E2F transcription factor do?
E2F transcription factors regulate the expression of a number of genes important in cell proliferation, particularly those involved in progression through G1 and into the S-phase of the cell cycle.
Is ef2 a transcription factor?
The E2 factor (E2F) family of transcription factors are downstream effectors of the retinoblastoma (RB) protein pathway and are believed to play a pivotal role in cell division control. Since its discovery, E2F has been viewed as a positive regulator of genes required for DNA synthesis.
What does E2F stand for?
E2FAcronymDefinitionE2FE-Mail to FaxE2FE2 Transcription Factor
Is E2F a promoter?
E2F promoter in pDRIVE expression plasmid E2F-1 is a transcription factor that activates its own transcription and that of other genes involved in the G(1) to S transition phase of the cell cycle.
Is E2F a tumor suppressor gene?
Although it can function as an oncogene, E2F-1 has also been postulated to be a tumor suppressor because it can induce apoptosis. Mice with a homozygous deletion of E2F-1 have a defect in thymocyte apoptosis, show increased cellular proliferation, and eventually develop numerous tumors [43,201].
What type of protein is E2F?
E2F is a group of genes that encodes a family of transcription factors (TF) in higher eukaryotes. Three of them are activators: E2F1, 2 and E2F3a. Six others act as suppressors: E2F3b, E2F4-8. All of them are involved in the cell cycle regulation and synthesis of DNA in mammalian cells.
What genes are regulated by E2F?
A role for E2F in the activation of several DNA replication genes has been well established. Known E2F targets include genes encoding proteins involved in the initiation of replication (Orc1, Mcm proteins, Cdc6), nucleotide metabolism (thymidine kinase), and the enzymatic synthesis of DNA (DNA polymerase α).
Is E2F an enzyme?
E2F-1 and a cyclin-like DNA repair enzyme, uracil-DNA glycosylase, provide evidence for an autoregulatory mechanism for transcription. J Biol Chem. 1995 Mar 10;270(10):5289-98.
What is Rb and E2F?
The RB/E2F pathway regulates apoptosis, and RB inhibition of apoptosis is an important mechanism of tumor suppression whereby cells deficient for RB function can be eliminated by apoptosis. One manner through which RB can inhibit apoptosis is through its binding to RNA processing factors.
Does E2F bind DNA?
The E2F domains of E2F4 and E2F8 bind to DNA in an almost identical manner, forming contacts with C5 C6 G8 of one strand and G7′ of the complementary strand, and in addition, form several backbone contacts with both strands.
How is E2F regulated?
The E2F proteins form a family of transcription factors that regulate the transition from the G1 to the S phase in the cell cycle. E2F activity is regulated by members of the retinoblastoma protein (pRb) family, ensuring the tight control of E2F-responsive genes.
How does Rb inhibit E2F?
Rb proteins are thought to inhibit expression of E2F-regulated genes in two ways (Dyson et al., 2002): by directly binding and blocking the activation domain of E2F proteins or by active repression through the recruitment of HDAC, SWI/SNF factors, Polycomb group proteins (Dahiya et al., 2001) or methyltransferase ( ...
Does E2F bind DNA?
The E2F domains of E2F4 and E2F8 bind to DNA in an almost identical manner, forming contacts with C5 C6 G8 of one strand and G7′ of the complementary strand, and in addition, form several backbone contacts with both strands.
What happens if E2F is inactivated?
The E2F transcription factors play a critical role in controlling cell fate. In Drosophila, the inactivation of E2F in either muscle or fat body results in lethality, suggesting an essential function for E2F in these tissues.
What are CDC6 and Cdt1?
CDC6 is an ATP binding protein and a member of the pre-replicative complex (pre-RC) together with the origin recognition complex (ORC), Cdt1 and the MCM complex (containing MCM2-7p). CDC6 assembles after ORC in an ATP dependent manner and is required for loading MCM proteins onto the DNA.
How does Rb inhibit E2F?
Rb proteins are thought to inhibit expression of E2F-regulated genes in two ways (Dyson et al., 2002): by directly binding and blocking the activation domain of E2F proteins or by active repression through the recruitment of HDAC, SWI/SNF factors, Polycomb group proteins (Dahiya et al., 2001) or methyltransferase ( ...
What are the targets of E2F transcription?
Among E2F transcriptional targets are cyclins, CDKs, checkpoints regulators, DNA repair and replication proteins. Nonetheless, there is a great deal of redundancy among the family members. Mouse embryos lacking E2F1, E2F2, and one of the E2F3 isoforms, can develop normally when either E2F3a or E2F3b, is expressed.
What is the role of E2F in the cell cycle?
E2F family members play a major role during the G1/S transition in mammalian and plant cell cycle (see KEGG cell cycle pathway ). DNA microarray analysis reveals unique sets of target promoters among E2F family members suggesting that each protein has a unique role in the cell cycle. Among E2F transcriptional targets are cyclins, CDKs, checkpoints regulators, DNA repair and replication proteins. Nonetheless, there is a great deal of redundancy among the family members. Mouse embryos lacking E2F1, E2F2, and one of the E2F3 isoforms, can develop normally when either E2F3a or E2F3b, is expressed.
What is the activator of E2F-3A?
The activation of E2F-3a genes follows upon the growth factor stimulation and the subsequent phosphorylation of the E2F inhibitor retinoblastoma protein, pR B. The phosphorylation of pRB is initiated by cyclin D / cdk4, cdk6 complex and continued by cyclin E/cdk2. Cyclin D/cdk4,6 itself is activated by the MAPK signaling pathway .
What are the two groups of E2F?
The E2F family is generally split by function into two groups: transcription activators and repressors. Activators such as E2F1, E2F2, E2F3a promote and help carryout the cell cycle, while repressors inhibit the cell cycle. Yet, both sets of E2F have similar domains.
What is the family of transcription factors?
Family of transcription factors. E2F is a group of genes that encodes a family of transcription factors (TF) in higher eukaryotes. Three of them are activators: E2F1, 2 and E2F3a. Six others act as suppressors: E2F3b, E2F4-8. All of them are involved in the cell cycle regulation and synthesis of DNA in mammalian cells.
How does pRb repress E2F-3A?
When bound to E2F-3a, pRb can directly repress E2F-3a target genes by recruiting chromatin remodeling complexes and histone modifying activities (e.g. histone deacetylase, HDAC) to the promoter.
Which family of transcription factors has a fold similar to the winged helix motif?
X-ray crystallographic analysis has shown that the E2F family of transcription factors has a fold similar to the winged-helix DNA-binding motif.
What is transcription factor E2F?
Transcriptional Regulation by the E2F Transcription Factors. E2F was originally identified as a cellular DNA binding activity that regulated expression of the viral E2 promoter.30,31 Since this seminal work, molecular analysis has revealed that the E2F activity is encoded by a family of DNA binding proteins, which includes transcriptional ...
What is the function of E2F?
E2F is a cellular transcription factor that is necessary for the transactivation of the adenoviral E2 promoter and thus the expression of viral DNA replication proteins.
What are the E2F family members?
The major member of the E2F family of transcription factors is E2F1 (DeGregori, 2002 ). Based on structure, transcriptional properties, and association with pocket proteins, the E2F family can be divided onto three distinct subgroups. The factors E2F1, E2F2, and E2F3 associate exclusively with pRb, are potent transcriptional activators, and can induce apoptosis. In contrast, E2F4, which associates with Rb, p107, and p130, and E2F5, which associates with p130, seem to be primarily involved in the active repression of E2F-responsive genes. Meanwhile, E2F6 does not interact with pocket proteins and functions as a negative regulator of E2F-dependent transcription via complexing with chromatin modifiers.
How many members are in the E2F family?
The E2F family of transcription factors consists of nine members, and these members are classically grouped as either activators (E2F1–3a) or repressors (E2F3b–8) (Chen et al., 2009). However, recent studies suggest that E2Fs can function as either activators or repressors of transcription depending on the cellular context, target genes, and cofactors involved (Chong et al., 2009; Lee, Bhinge, & Iyer, 2011 ). The classical E2Fs (E2F1–6) contain one DNA-binding domain and form heterodimers with DP proteins. The more recently identified members, E2F7 and E2F8, referred to as atypical E2Fs as they possess two DNA-binding domains, form homo- or heterodimers, and regulate transcription in a DP-independent manner ( Lammens et al., 2009 ). Although deletion of E2F7/8 in mice resulted in embryonic death by day E11.5, they did not show any defects in proliferation ( Li et al., 2008); instead, these mice displayed massive apoptosis and vascular defects at E10.5. Interestingly, apoptosis, but not vascular defects, were rescued by additional deletion of E2F1 or p53 in these double knock-out mice, indicating that E2F7/8 regulate vascular integrity through an alternate mechanism (Li et al., 2008 ). Further studies revealed that E2F7 and E2F8 could directly bind to the promoter of VEGFA and activate transcription, independent of canonical E2F-binding elements ( Bakker, Weijts, Westendorp, & de Bruin, 2013; Weijts et al., 2012). Instead, E2F7/8 form a transcriptional complex with HIF-1 to stimulate VEGFA promoter induction. These results uncover an unexpected link between atypical E2Fs and HIF1–VEGFA pathway, adding to the complexity of the molecular mechanisms by which Rb–E2F pathway regulates angiogenesis.
What are the roles of E2F and DP in the cell cycle?
The E2F, DP, and Rb families play key roles in regulation of the mammalian cell cycle. E2F and DP family members comprise transcription factors that regulate transcription of genes encoding proteins necessary for progression through S phase of the cell cycle, for example, DHFR, Orcl, Myb, Myc, cdc2, and others (56, 57). The E2F and Rb genes are expressed in a cell cycle-dependent manner. E2F family members bind E2F sites as heterodimers with members of the DP family (56, 57). E2F/DP heterodimers can be negatively regulated by association with the Rb family of proteins, including Rb (p110), p107, and p130 (58). Rb is frequently mutated in human cancer, reflecting the importance of regulation of E2F/DP and Rb family members in cell cycle progression. From the immediate perspective, E2F1, DP, and Rb families are of further interest because their regulation is dependent on the reciprocal induction and inhibition of cell cycle-specific transcription from their own genes. Not all of these factors are strictly autoregulatory, but instead it is the families of genes that are autoregulated.
What is the 106b25 gene?
The 106b~25 polycistronic cluster, which promoted bypass of RIS ( Borgdorff et al., 2010 ), is contained within an intronic region of the MCM7 gene . MCM7 is part of a multiprotein helicase complex required for DNA replication and G 1 /S transition, and represents a marker of high proliferation rate in cancer cells ( Blow and Hodgson, 2002 ). In gastric primary tumors, all three members of the 106b~25 cluster, MiR-106b, -93, and -25, were coregulated with MCM7. Furthermore, MiR-106b and -93 were shown to directly target E2F1 mRNA, while blocking TGFβ tumor suppressor activity via direct inhibition of its downstream effector CDKN1A ( Petrocca et al., 2008 ). E2F1 displays oncogenic properties through its major role as a regulator of G1/S transition; however, unlike other E2F members, its accumulation beyond a threshold triggers an intrinsic tumor suppressor activity inducing apoptosis both in vitro and in vivo ( Lazzerini Denchi and Helin, 2005 ). These observations suggest that downregulation of E2F1 by the 106b~25 cluster participates in tumor proliferation in gastric cancer.
What are the genes that are regulated by E2F?
In this review the genes known to be regulated by E2F are sorted into four general groups. The first group includes genes whose protein products are necessary for DNA replication. This includes genes encoding ORC and MCM proteins that are needed for the initiation of DNA synthesis, and genes encoding proteins such as dihydrofolate reductase and ribonucleotide reductase that are needed for the enzymatic synthesis of DNA. A second group of genes contains critical cell cycle regulators such as cyclin A, cyclin E, and cdc25c that act to drive cell cycle progression. Because E2F-binding sites are found in the promoters of so many essential genes, it is thought that E2F and pRB exert control of the G1/S transition by coordinately controlling the expression of large programs of transcription rather than by acting on a single essential target.
What is E2F transcription?
The E2 factor (E2F) family of transcription factors are downstream targets of the retinoblastoma protein. E2F factors have been known for several years to be important regulators of S-phase entry. Recent studies have improved our understanding of the molecular mechanisms of action used by this transcriptional network. In addition, they have given us an appreciation of the fact that E2F has functions that reach beyond G1/S control and impact cell proliferation in several different ways. The discovery of new family members with unusual properties, the unexpected phenotypes of mutant animals, a diverse collection of biological activities, a large number of new putative target genes and the new modes of transcriptional regulation have all contributed to an increasingly complex view of E2F function. In this review, we will discuss these recent developments and describe how they are beginning to shape a new and revised picture of the E2F transcriptional program.
What is the E2F gene?
Since the initial identification of the cellular factor required for the activation of the E2 viral promoter, the E2 factor (E2F), (reviewed in Dyson, 1998; Nevins, 1998 ), the E2F family of transcription factors has been growing steadily through the addition of new members in mammals and through the discovery of homologs in other eukaryotes. This expansion has led to an appreciation of the differences between E2F members. Individual E2Fs have distinct mechanisms of action, and regulation, and are associated with particular types of biological activities.
How does E2F affect the cell cycle?
Studies in both mammalian cells and in worms point to a role for E2F in the inhibition of the anaphase-promoting complex, and this may be another means by which E2F influences cell cycle progression ( Lukas et al., 1999; Fay et al., 2002; Garbe et al., 2004 ). Saavedra and colleagues reported that inactivation of E2F3 leads to centrosome amplification, mitotic spindle defects and aneuploidy, and suggested that E2F3 functions to coordinate the DNA replication with the centrosome duplication cycle ( Saavedra et al., 2003 ). Work in Drosophila suggests that E2F is an important determinant of the length of time that cells spend in different phases of the cell cycle. Reis and Edgar proposed that E2F proteins, by acting on both the G1/S and G2/M transitions, enable cells to maintain normal proliferation rates by altering the length of G2 in response to alterations in the length of G1 ( Reis and Edgar, 2004 ). Perhaps, the clearest evidence that E2F is generally required for cell cycle progression, rather than acting solely at the G1/S transition, comes from the phenotype observed in MEFs when all three activator E2Fs were inactivated. E2f1−/−; E2f2−/−; E2f3−/− cells do not arrest in G1 but appear to be blocked at all phases of the cell cycle ( Wu et al., 2001 ), suggesting that E2F is needed to drive cells through multiple cell cycle transitions.
How does E2F1-5 affect transcription?
These interactions negatively affect E2F-dependent transcription in two ways – binding to E2Fs inhibits their ability to activate transcription. In addition, when pRB-related proteins are recruited to promoters by repressor E2Fs, they can actively repress transcription by associating with various chromatin-modifying repressive complexes (reviewed in Harbour and Dean, 2000a; Ferreira et al., 2001; Frolov and Dyson, 2004 ). E2F activity is also regulated by various additional mechanisms (for a detailed review see Helin, 1998; Muller and Helin, 2000 ).
Why is it so difficult to determine how E2F activates and RB family members repress transcription?
Why has it been so difficult to determine how E2F activates and RB family members repress transcription? One reason may be that many studies have focused on mechanisms of repression used by pRB at ‘classic’ E2F target genes. The problem here is that it is far from certain that pRB is directly required for the regulation of these targets for much of the time. Many cell cycle-regulated E2F targets are derepressed in p107−/−;p130−/− but show little or no changes in Rb−/− cells ( Herrera et al., 1996; Hurford et al., 1997; Mulligan et al., 1998 ). Several studies have failed to detect pRB at these promoters, whereas p107 and p130 are readily detectable ( Takahashi et al., 2000; Wells et al., 2000; Aslanian et al., 2004 ). Indeed, the analysis of Rb−/− cells revealed that pRB is not required for HDAC recruitment to several of the traditional E2F targets ( Rayman et al., 2002 ). Thus, either pRB does not regulate these promoters directly or it may be recruited there only under specific circumstances, as has been proposed ( Dahiya et al., 2001; Narita et al., 2003 ). It is uncertain whether all pocket proteins use the same mechanisms of repression and it may not be accurate to extend conclusions drawn from studies of one family member to the others.
Why is E2F important?
The idea that E2F/RB proteins are important during development is further supported by studies in flies, worms and frogs. In C. elegans E2F is required for embryonic asymmetry ( Page et al., 2001 ), and E2F and RB antagonize Ras signaling during vulval development ( Ceol and Horvitz, 2001 ). In Drosophila, E2F is required for the establishment of the dorsoventral polarity in the oocyte ( Myster et al., 2000; Cayirlioglu et al., 2001 ), whereas in Xenopus, E2F is required for ventral and posterior cell fate determination during early embryogenesis ( Suzuki and Hemmati-Brivanlou, 2000 ).
What is E2F1 bound to?
For instance, E2F1 is bound to the caspase-7 promoter in cells expressing the E1A viral oncogene ( Nahle et al., 2002 ). Similarly, E2F1 is recruited to the Arf promoter only in response to various oncogenic stimuli ( Aslanian et al., 2004; PJ Iaquinta and JA Lees, personal communication). At the Apaf1 promoter, expression is induced and E2F bound only when E2F1 activity is elevated ( Moroni et al., 2001; Furukawa et al., 2002 ). These responses are not unique to E2Fs. It has been proposed that pRB is recruited to cell cycle promoters only during senescence ( Dahiya et al., 2001; Narita et al., 2003 ), and p107 containing complexes are specifically recruited to the c-myc promoter upon TGF β signaling ( Chen et al., 2002 ).

Overview
E2F/pRb complexes
The Rb tumor suppressor protein (pRb) binds to the E2F1 transcription factor preventing it from interacting with the cell's transcription machinery. In the absence of pRb, E2F1 (along with its binding partner DP1) mediates the trans-activation of E2F1 target genes that facilitate the G1/S transition and S-phase. E2F targets genes that encode proteins involved in DNA replication (for example DNA polymerase, thymidine kinase, dihydrofolate reductase and cdc6), and chromosoma…
Genes
Homo sapiens E2F1 mRNA or E2F1 protein sequences from NCBI protein and nucleotide database.
Structure
X-ray crystallographic analysis has shown that the E2F family of transcription factors has a fold similar to the winged-helix DNA-binding motif.
Role in the cell cycle
E2F family members play a major role during the G1/S transition in mammalian and plant cell cycle (see KEGG cell cycle pathway). DNA microarray analysis reveals unique sets of target promoters among E2F family members suggesting that each protein has a unique role in the cell cycle. Among E2F transcriptional targets are cyclins, CDKs, checkpoints regulators, DNA repair and replication pro…
Transcriptional targets
• Cell cycle: CCNA1,2, CCND1,2, CDK2, MYB, E2F1,2,3, TFDP1, CDC25A
• Negative regulators: E2F7, RB1, TP107, TP21
• Checkpoints: TP53, BRCA1,2, BUB1
• Apoptosis: TP73, APAF1, CASP3,7,8, MAP3K5,14
See also
• Transcription factor DP
• Type 3c (Pancreatogenic) Diabetes
External links
• E2F+Transcription+Factors at the US National Library of Medicine Medical Subject Headings (MeSH)
• Drosophila E2F transcription factor - The Interactive Fly
• Drosophila E2F transcription factor 2 - The Interactive Fly