
Pseudogene
Pseudogenes are functionless relatives of genes that have lost their gene expression in the cell or their ability to code protein. Pseudogenes often result from the accumulation of multiple mutations within a gene whose product is not required for the survival of the organism.
What is a pseudogene?
A pseudogene is a DNA sequence that resembles a gene but has been mutated into an inactive form over the course of evolution. A pseudogene shares an evolutionary history with a functional gene and can provide insight into their shared ancestry.
What is the function of pseudogenes in Nos gene expression?
The actual pseudogene function that regulates NOS gene expression is believed to operate as follows: ‘Specifically, the active transcription of the pseudogene will lead to the suppression of nNOS protein synthesis, and on the other hand, the inhibition of pseudogene transcription will permit nNOS production.
What is the difference between functional and pseudogenes?
Pseudogenes are inheritable genetic elements that are similar to functional genes but are non-functional as they do not encode for proteins. Their biogenesis results from the duplication of a parental gene, or the retrotransposition of an mRNA sequence into different genomic loci.
What is the role of structural information in the study of pseudogenes?
In the case of pseudogenes, structural information can give extra evolutionary clues and facilitate analysis of the scope of folds in the pseudogene population ("pseudo"-folds) in contrast to those observed for the genes themselves.
What defines a pseudogene?
Listen to pronunciation. (SOO-doh-jeen) A DNA sequence that resembles a gene but has been mutated into an inactive form over the course of evolution. It often lacks introns and other essential DNA sequences necessary for function.
What makes a gene a pseudogene?
Definition. A pseudogene is a segment of DNA that structurally resembles a gene but is not capable of coding for a protein. Pseudogenes are most often derived from genes that have lost their protein-coding ability due to accumulated mutations that have occurred over the course of evolution.
What is the difference between a pseudogene and a gene?
A pseudogene is an inheritable genetic element that is nonfunctional as it does not code for a protein. On the other hand, a gene is an inheritable genetic element that is functional as it does code for a specific protein. So, this is the key difference between pseudogene and gene.
What is an example of a pseudogene?
Pseudogenes are alleles of normal genes that have become non-functional due to accumulation of mutations; for example, the protein coding region may contain a premature stop codon, or a frameshift mutation, or an internal deletion or insertion relative to the normal sequence.
What is a pseudogene quizlet?
What is a pseudogene? previously functional gene that lost its function due to mutation.
What is pseudogenes and its types?
Pseudogenes come in two basic types according to their precise mode of origin: processed pseudogenes that lack introns because they arise when functional messenger RNA is retrotranspositionally inserted into the genome; and non-processed pseudogenes that are the evolutionary remnants of tandemly duplicated genes that ...
Why do organisms keep pseudogenes?
Pseudogenes play an essential role in comparative studies regarding genomics as they can provide a record of ancient genes. They are used to determine the rate of gene duplication and follow the evolution of sequence changes in organisms. Thus, pseudogenes are unique and helpful for phylogenetic studies.
Why are pseudogenes non functional?
Pseudogenes are inheritable genetic elements that are similar to functional genes but are non-functional as they do not encode for proteins. Their biogenesis results from the duplication of a parental gene, or the retrotransposition of an mRNA sequence into different genomic loci.
What pseudogenes do humans have?
We identified ∼20,000 pseudogenes in the human genome. The strategy used in this study ensures that each pseudogenic region represents a single event of gene or exon duplication and that regions matching to the same protein are fused.
How do you identify pseudogenes?
All of them identify pseudogenes based on their two key sequence properties: similarity to genes and non-functionality. In practice, the former is often characterized by the sequence similarity between a pseudogene and its closest functioning gene relative (referred to as the 'parent gene') in the present-day genome.
How pseudogenes are different from noncoding RNA genes?
Noncoding RNAs Pseudogenes are inheritable genetic elements that are similar to functional genes but are non-functional as they do not encode for proteins. Their biogenesis results from the duplication of a parental gene, or the retrotransposition of an mRNA sequence into different genomic loci.
How can gene duplications occur?
Gene duplication can occur as the result of an error in recombination or through a retrotransposition event. Duplicate genes are often immune to the selective pressure under which genes normally exist. This can result in a large number of mutations accumulating in the duplicate gene code.
Why do organisms keep pseudogenes?
Pseudogenes play an essential role in comparative studies regarding genomics as they can provide a record of ancient genes. They are used to determine the rate of gene duplication and follow the evolution of sequence changes in organisms. Thus, pseudogenes are unique and helpful for phylogenetic studies.
Which is an example of Subfunctionalization of a gene duplicate?
Hemoglobin. Human hemoglobin provides a variety of subfunctionalization examples. For instance, the gene for hemoglobin α-chain is undoubtedly derived from a duplicate copy of hemoglobin β-chain.
What is pseudogenes in biology?
For a species of beetle, see Pseudogenes (beetle). Pseudogenes are nonfunctional segments of DNA that resemble functional genes. Most arise as superfluous copies of functional genes, either directly by DNA duplication or indirectly by reverse transcription of an mRNA transcript.
What are the properties of pseudogenes?
Properties. Pseudogenes are usually characterized by a combination of homology to a known gene and loss of some functionality. That is, although every pseudogene has a DNA sequence that is similar to some functional gene, they are usually unable to produce functional final protein products.
What is the pseudogene of Drosophila?
The term "pseudo-pseudogene" was coined for the gene encoding the chemosensory ionotropic glutamate receptor Ir75a of Drosophila sechellia, which bears a premature termination codon (PTC) and was thus classified as a pseudogene.
How to identify pseudogenes?
Pseudogenes are often identified by the appearance of a premature stop codon in a predicted mRNA sequence , which would, in theory, prevent synthesis ( translation) of the normal protein product of the original gene. There have been some reports of translational readthrough of such premature stop codons in mammals. As alluded to in the figure above, a small amount of the protein product of such readthrough may still be recognizable and function at some level. If so, the pseudogene can be subject to natural selection. That appears to have happened during the evolution of Drosophila species .
What is the PTEN gene?
PTEN. The PTEN gene is a known tumor suppressor gene. The PTEN pseudogene, PTENP1 is a processed pseudogene that is very similar in its genetic sequence to the wild-type gene.
What are unitary pseudogenes?
Unitary pseudogenes. 2 ways a pseudogene may be produced. Various mutations (such as indels and nonsense mutations) can prevent a gene from being normally transcribed or translated, and thus the gene may become less- or non-functional or "deactivated".
What happens when a gene is mutated?
Mutations that disrupt either the structure or the function of either of the two genes are not deleterious and will not be removed through the selection process. As a result, the gene that has been mutated gradually becomes a pseudogene and will be either unexpressed or functionless.
Summary
The discovery of a functional nitric oxide synthase (NOS) pseudogene compels us to understand pseudogenes in a new light. It confirms earlier clues suggesting that seemingly nonfunctional pseudogenes can regulate the expression of paralogous genes by producing antisense RNA.
Potential modes of pseudogene function
It has been demonstrated that pseudogenic features, notably seemingly absent or disabled promoters, premature stop codons, splicing errors, frameshift-causing deletions and insertions, etc., do not necessarily abolish gene expression.
Voices crying out in the wilderness
Against the backdrop of the customary negative opinion of pseudogenes, there have always been a few individuals who anticipated their functional potential. McCarrey et al. 8 were probably the first to suggest that pseudogenes can be functional in terms of the regulation of the expression of its paralogous genes.
A functional antisense RNA-producing pseudogene
In the snail Lymnea stagnalis, the neuronal enzyme NO synthase (nNOS) is encoded by the NOS gene (now called the Lym -nNOS gene). The enzyme induces the production of nitrogen oxide (NO), an intracellular signaling molecule in the snail’s nervous system. One function of (NO) is the mediating of its feeding behavior.
Protein suppression by a second functional antisense pseudogene?
A second functional (pseudo)gene, occurring in tandem with the first one (see Figure 2) is now known to exist:
Functional mildly-conserved pseudogene nucleotide sequences
A long held ostensible support for the absence of pseudogene function has been their usual apparent lack of sequence conservation. Protein-coding genes typically vary only slightly among orthologs and paralogs as a result of purifying selection*.
Multiple modes of pseudogenic antisense RNA production
In conventional protein synthesis, the process begins with the transcription of the DNA molecule in a 5’ to 3’ (left to right) direction, and the complementary DNA strand is not used.
What are pseudogenes in the genome?
Pseudogenes are regions of the genome that are similar to functional genes but are thought to be nonfunctional. They may be highly homologous to protein-coding genes but unable to produce a functional protein due to a disrupted open reading frame (ORF) or highly similar to RNA encoding genes but unable to produce an RNA transcript. RNA pseudogenes are more difficult to identify as there is no ‘ORF’ to be disrupted; nevertheless, the term ‘pseudogene’ was first used by Jacq et al. (1977) to describe RNA pseudogenes of the 5S RNA gene that are found in a tandem array with the functional 5S RNA gene in Xenopus. Pseudogenes have often been regarded as the ‘poor relations’ of protein-coding genes and have elicited little interest from researchers aside from their potential to elucidate the evolutionary processes that have been acting in the genome. However, some recent studies have suggested that pseudogenes have the potential to act as posttranscriptional regulators of functional genes, by both encoding short interfering RNAs and acting as sinks for ncRNAs, in particular miRNAs ( Muro et al., 2011 ). Further, it is possible for pseudogenes to be involved in gene conversion with functional genes, resulting in disease phenotypes ( Bischof et al., 2006 ), and for pseudogenized loci to be resurrected, possibly again by gene conversion ( Wang et al., 2012 ). It is also critical that pseudogenes are correctly annotated within genomes to avoid confusing them with coding loci. Several groups have attempted to catalog all pseudogenes within the human genome; one of the most recent is the GENCODE gene annotation project of the ENCODE consortium, which has annotated over 12 000 human pseudogenes that are stored at the psiDR website ( www.pseudogenes.org/psidr/) ( Pei et al., 2012 ).
How are pseudogenes related to functional genes?
Pseudogenes are inheritable genetic elements that are similar to functional genes but are non-functional as they do not encode for proteins. Their biogenesis results from the duplication of a parental gene, or the retrotransposition of an mRNA sequence into different genomic loci. The inability of pseudogenes to produce functional proteins is often the consequence of subsequent genetic alterations (frameshift mutations, creation of stop codon). There exist roughly 10,000 pseudogenes in mammalian genomes. Although they are not able to produce functional proteins, many pseudogenes (approximately 20%) are transcribed into RNAs that comprise another category of lncRNAs [33]. Several studies demonstrated the biological activity of pseudogenes in controlling parental gene expression by producing natural siRNAs [34] or antisense transcripts, and by competing with miRNA binding sites on mRNA targets [35].
What is the pseudogene of D. melanogaster?
A different type of pseudogene, known as a processed pseudogene, is found in the GST supergene family in D. melanogaster. Mgst1-psi is a putative pseudogene of the single microsomal GST gene found in this species. This pseudogene contains a poly-A tail towards the 3′ end, has no introns, a premature stop codon, and a direct repeat at both ends. Presumably, it was derived from the reverse transcription and subsequent integration of Mgst1 mRNA into the genome ( Toba and Aigaki, 2000 ).
How many pseudogenes are there in the human genome?
There exist roughly 10,000 pseudogenes in mammalian genomes.
How are pseudogenes produced?
Pseudogenes can be produced by deleterious mutations, which result in the silencing of a gene. This type of pseudogene, known as an unprocessed pseudogene, normally occurs in a duplicate gene and frequently pseudogenes are found proximal to the genes from which they are derived.
What is a unitary pseudogene?
A well-known example of a unitary pseudogene in the human genome is the GULOP locus, which is a pseudogenized version of the gene encoding gulonolactone (L-) oxidase that processes ascorbic acid (vitamin C) and is functional (GULO) in most other vertebrates ( Zhu et al., 2007 ).
Which miRNAs regulate PTEN?
The studies suggest that PTEN is regulated by several miRNAs viz., miR-17, miR-19, miR-21, miR-26, and miR-214; and PTENP1 acts as a decoy to derepress the inhibitory activity of these miRNAs [92,286,287].
What is pseudogene nonfunctionality?
Pseudogene nonfunctionality tacitly assumes that any functional peptide synthesized should be the same or very similar to that encoded by the paralogous protein-coding gene. The actual or perceived inability of the pseudogene to direct synthesis of such a peptide is conventionally taken as proof of its ‘junk’ status. However, this long-held premise can no longer be sustained. It is now known that a snail’s pseudogene can direct the synthesis of a useful shortened peptide. This truncated peptide can form a complex with the full-length peptide produced by the paralogous gene, thus functioning as a regulator of the abundance of this full-length protein. 5
Is Makorin1-P1 a paralog?
It is commonly supposed that a pseudogene lacks function, relative to its counterpart gene paralog, because it lacks an entire large segment of sequence. The Makorin1-p1 pseudogene demonstrates that this is not the case. Only the first 700 nucleotides of the mRNA transcript of this pseudogene correspond approximately to the mRNA transcript of the paralogous Makorin1 gene. Yet the fragmentary pseudogene DNA segment that is transcribed into this RNA sequence is more than sufficient for the Makorin1-p1 pseudogene to perform its function. This pointedly warns against assuming that even a highly fragmented pseudogene inevitably lacks function.
What is pseudogenes in biology?
Pseudogenes are genomic DNA sequences similar to normal genes but non-functional; they are regarded as defunct relatives of functional genes. What causes pseudogenes to arise? There are two accepted processes during which pseudogenes may arise: duplication - modifications (mutations, insertions, deletions, frame shifts) to the DNA sequence ...
Why is structural information important in the case of pseudogenes?
In the case of pseudogenes, structural information can give extra evolutionary clues and facilitate analysis of the scope of folds in the pseudogene population ("pseudo" -folds) in contrast to those observed for the genes themselves.
What is the name of the program that detects a gene homologous region?
Once gene sequences have been identified in the genome, it is possible to use sequence alignment programs (such as FASTA or BLAST) to detect matching regions in the nucleotide sequence. These matching regions are potential gene homologs and are termed pseudogenes if there is some evidence that either of the causes (see above) are satisfied.
What is a copy of a gene that is disabled?
Copies of genes that are disabled in such a manner are termed non-processed or duplicated pseudogenes. retrotransposition - reverse transcription of an mRNA transcript with subsequent re-integration of the cDNA into the genome. Such copies of genes are termed processed pseudogenes.
How can evolutionary relationships between proteins be elucidated?
In a number of cases, more distant evolutionary and functional relationships between proteins can only be elucidated through the analysis of the folds that their structures adopt. While it must not be forgotten that the assignment of function to a gene is often implied from that of a gene with a homologous sequence, the added information that protein structures can provide is very desirable in genome annotation.
What is the goal of the eukaryotic genome survey?
Our initial goal was to survey some eukaryotic genomes for pseudogene sequences and fragments of pseudogene sequences. In addition to this, we have also quantified "pseudo-fold" usage, amino-acid composition, and single-nucleotide polymorphisms (SNPs) to help elucidate the relationships between pseudogene families across these organisms.
How do pseudogenes function?
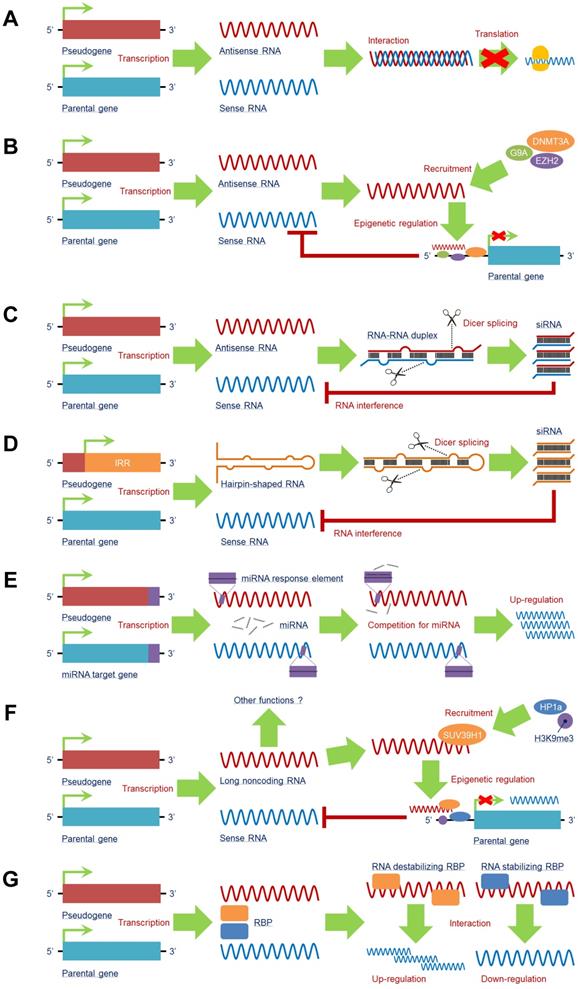
Another mechanism through which pseudogenes can function is by influencing chromatin or genomic architecture (Fig. 2e ). HBBP1, a pseudogene residing within the haemoglobin locus, enables the dynamic chromatin changes that regulate expression of fetal and adult globin genes during development 50. Notably, although inhibiting HBBP1 transcription has no effect, deletion of the genomic locus reactivates fetal globin expression. HBBP1 DNA contacts, but not transcription, are required for suppressing the expression of fetal globin genes in adult erythroid cells 50. Another example in which pseudogenes may function intrinsically as DNA elements is by influencing chromosome stability. A study of the genomic architecture of the 22q11.2 locus, which is associated with DiGeorge/velocardiofacial syndrome, proposed that deletions of a pseudogene within the low-copy repeat region could increase non-allelic homologous recombination, which in turn results in deletions that underlie the disease 51. Pseudogenes can also produce pseudogene–gene fusion transcripts. In prostate cancer, exons of the pseudogene KLKP1 are spliced into the adjacent KLK4 gene, creating a novel fusion protein 52, 53. Polymorphic retrocopies can also form fusion transcripts if they are located in the intron of a gene. For example, a polymorphic retrocopy of CBX3 located in an intron of CCDC32 results in a chimeric transcript of unknown function in some individuals 15. Pseudogenes can also transfer deleterious alleles to their parental genes by non-allelic recombination (gene conversion) 54 (Fig. 2f ). Pseudogene-mediated gene conversion underlies cases of hereditary pancreatitis 55, adrenal hyperplasia 56, polycystic kidney disease 57, cataracts 58 and a multitude of other diseases 54.
What is a pseudogene?
Pseudogenes are defined as regions of the genome that contain defective copies of genes. They exist across almost all forms of life, and in mammalian genomes are annotated in similar numbers to recognized protein-coding genes. Although often presumed to lack function, growing numbers of pseudogenes are being found to play important biological roles.
What is pseudogene annotation?
Annotations of metazoan genomes typically describe between 10,000 and 20,000 regions as pseudogenic 16. The binary distinction between genes and pseudogenes forms a central theme in genome annotation and, ultimately, informs the reference list of genes of an organism. Consequently, the annotation of genomic regions as pseudogenes constitutes an etymological signifier that an element has no function and is not a gene. As a result, pseudogene-annotated regions are largely excluded from functional screens and genomic analyses 17, 18, 19. Therefore, the process of pseudogene annotation is paramount in the consideration of which genomic elements are assessed for biological impact. However, with a growing number of instances of pseudogene-annotated regions later found to exhibit biological function, there is an emerging risk that these regions of the genome are prematurely dismissed as pseudogenic and therefore regarded as void of function. Due to the recent maturation of several enabling technologies, we propose that the time is opportune for a re-evaluation of the functions of pseudogene-annotated regions. The advent of long-read transcriptomics enables the identification of dynamically active pseudogenes, and whole-genome sequencing of large cohorts enables the identification of disease-associated pseudogenes. Additionally, the CRISPR–Cas9 revolution allows straightforward interrogation of pseudogene functions.
What are the processes of gene duplication and retrotransposition?
Gene duplication and retrotransposition are crucial processes that underlie organismal robustness and the evolution of new biological functions and characteristics. These processes sit alongside other evolutionary mechanisms, such as meiosis, that create the organismal diversity upon which natural selection acts. Like other evolutionary processes, gene duplication and retrotransposition leave behind observable ancestral signatures that provide insight into an organism’s history. In the fundamental reductionist approach often assumed in genetics and molecular biology, the perspective is often lost that life as we observe it today is not only the product of billions of years of evolutionary processes but also still subject to these same processes. Although the pseudogene concept arose to describe an individual molecular phenomenon, the term was rapidly adopted to annotate tens of thousands of genomic regions that met only loosely defined criteria and was effectively axiomatized without being subject to any rigorous scientific debate. This lack of consensus-seeking process has left genome biology with a legacy concept that obscures objective investigation of genome function.
How to identify pseudogenes in eukaryotic genome?
Pseudogenes in eukaryotic genomes are identified by computational pipelines and manual annotation 5. Pseudogenes are first identified by searches for sequences similar to known genes 22, 23. The absence of introns, the occurrence of truncations and disruptions to the open reading frame relative to the parent gene are the primary characteristics used to identify pseudogenes 20. Different laboratory groups use various combinations of characteristics to identify putative pseudogenes 20, 24. Importantly, the absence of introns and the absence of strong evidence of transcription are sufficient to identify a processed pseudogene. Additionally, some processed pseudogenes do not harbour truncating mutations and have the same protein-coding capacity as their parent genes 8, 25. Indeed, 8.9% of recognized human protein-coding genes do not contain introns and are likely to be derived from retrotransposition 26. Thus, computational differentiation of pseudogenes from genes on a purely rule-based system is unlikely to be feasible as it will inherently conflict with many protein-coding genes derived through gene duplication and retrotransposition. Accordingly, untruncated pseudogenes are often manually reannotated as protein-coding genes if they have demonstrated function (for example, PGK2, POU5F1B and NANOGP8) 5.
Why is pseudogenes used in biology?
In the case of pseudogenes, the original use of the term rapidly proliferated to encompass a heterogeneous class of genomic elements. Despite early attempts to provide a systematic and scientifically grounded nomenclature to these genomic elements 125, the term ‘pseudogene’ became widely used to describe a potentially defective copy of a gene, and pseudogene identification became a routine process in the annotation of genomes.
Why can't DNA oligonucleotides differentiate from its parent?
DNA oligonucleotides are typically unable to differentiate a pseudogene from its parental gene (or other pseudogene copies) due to sequence similarity 24, 89 , precluding unambiguous identification of their transcript levels (Fig. 3a ). The expression dynamics and specificity of pseudogenes remain largely unknown.

Overview
Examples of pseudogene function
While the vast majority of pseudogenes have lost their function, some cases have emerged in which a pseudogene either re-gained its original or a similar function or evolved a new function. Examples include the following:
Drosophila glutamate receptor. The term "pseudo-pseudogene" was coined for the gene encoding the chemosensory ionotropic glutamate receptor Ir75a of Dr…
Properties
Pseudogenes are usually characterized by a combination of homology to a known gene and loss of some functionality. That is, although every pseudogene has a DNA sequence that is similar to some functional gene, they are usually unable to produce functional final protein products. Pseudogenes are sometimes difficult to identify and characterize in genomes, because the two requirements of homology and loss of functionality are usually implied through sequence align…
Types and origin
There are four main types of pseudogenes, all with distinct mechanisms of origin and characteristic features. The classifications of pseudogenes are as follows:
In higher eukaryotes, particularly mammals, retrotransposition is a fairly common event that has had a huge impact on the composition of the genome. For exa…
Misidentified pseudogenes
Sometimes genes are thought to be pseudogenes, usually based on bioinformatic analysis, but then turn out to be functional genes. Examples include the Drosophila jingwei gene which encodes a functional alcohol dehydrogenase enzyme in vivo.
Another example is the human gene encoding phosphoglycerate mutase which was thought to be a pseudogene but which turned out to be a functional gene, now named PGAM4. Mutations in it …
Bacterial pseudogenes
Pseudogenes are found in bacteria. Most are found in bacteria that are not free-living; that is, they are either symbionts or obligate intracellular parasites. Thus, they do not require many genes that are needed by free-living bacteria, such as gene associated with metabolism and DNA repair. However, there is not an order to which functional genes are lost first. For example, the oldest pseudoge…
See also
• List of disabled human pseudogenes
• Molecular evolution
• Molecular paleontology
• Pseudogene (database)
Further reading
• Gerstein M, Zheng D (August 2006). "The real life of pseudogenes". Scientific American. 295 (2): 48–55. Bibcode:2006SciAm.295b..48G. doi:10.1038/scientificamerican0806-48. PMID 16866288.
• Torrents D, Suyama M, Zdobnov E, Bork P (December 2003). "A genome-wide survey of human pseudogenes". Genome Research. 13 (12): 2559–67. doi:10.1101/gr.1455503. PMC 403797. PMID 14656963.
Summary
- More and more noncoding DNA, long considered ‘junk DNA’, has eventually been found to be functional.1–3 Hardly more than a few months pass by and there is not another scientific paper demonstrating function for some form of junk DNA. As summarized in this article, there is also growing evidence that at least some pseudogenes are functional. It should be stressed that pse…
Potential Modes of Pseudogene Function
- It has been demonstrated that pseudogenic features, notably seemingly absent or disabled promoters, premature stop codons, splicing errors, frameshift-causing deletions and insertions, etc., do not necessarily abolish gene expression.6 In fact, it is astonishing to realize that so-called pseudogenic features, instead of being ‘gene killing’ mutatio...
Voices Crying Out in The Wilderness
- Against the backdrop of the customary negative opinion of pseudogenes, there have always been a few individuals who anticipated their functional potential. McCarrey et al.8were probably the first to suggest that pseudogenes can be functional in terms of the regulation of the expression of its paralogous genes. They noted that the sense RNA transcribed* by a gene could be effectivel…
A Functional Antisense Rna-Producing Pseudogene
- In the snail Lymnea stagnalis, the neuronal enzyme NO synthase (nNOS) is encoded by the NOS gene (now called the Lym-nNOS gene). The enzyme induces the production of nitrogen oxide (NO), an intracellular signaling molecule in the snail’s nervous system. One function of (NO) is the mediating of its feeding behavior. Korneev, Park, and O’Shea14 were probably the first to provid…
Protein Suppression by A Second Functional Antisense Pseudogene?
- A second functional (pseudo)gene, occurring in tandem with the first one (see Figure 2) is now known to exist: This peptide, however, lacks certain functional domains.21Ordinarily, this would be taken as an obvious indicator of the fact that the protein is fatally defective and thus devoid of function, as is the pseudogene that directs its synthesis. Counter intuitively, however, this protei…
Functional Mildly-Conserved Pseudogene Nucleotide Sequences
- A long held ostensible support for the absence of pseudogene function has been their usual apparent lack of sequence conservation. Protein-coding genes typically vary only slightly among orthologs and paralogs as a result of purifying selection*. This is a result of the fact that most proteins cannot tolerate more than a few alterations without a marked detriment to their functio…
Multiple Modes of pseudogenic Antisense RNA Production
- In conventional protein synthesis, the process begins with the transcription of the DNA molecule in a 5’ to 3’ (left to right) direction, and the complementary DNA strand is not used. When antisense-mediated pseudogenic regulation of genes, by pseudogenes, was first proposed, the antisense RNA was envisioned as originating from a 5’ to 3’ transcription of the anticoding stran…
The Created Origin of The Functional Pseudogenes
- Let us evaluate the above-discussed evolutionary scenario (Figure 2) for the origin of these functional pseudogenes. To begin with, the relatively low degree of sequence similarity between the paralogous gene and pseudogenes weakens the argument that they necessarily arose from a common ancestral gene. Most re-arrangements of DNA segments within functional genes are h…
Conclusion
- Exciting new evidence is now surfacing on the functionality of pseudogenes. Korneev et al.assess the significance of their discovery as follows: No longer can it be assumed (contra Max4) that pseudogenes are just useless evolutionary discards. In fact, Korneev et al.’s21discovery prompts them to suggest that theirs is but the first discovery of an entirely new class of regulatory gene. …
Glossary
- Antisense RNA—An RNA molecule that, for whatever reason, is transcribed in a backwards (that is, tail to head, or 3’ to 5’) direction. Complementarity—The pairing off of nucleotides, between a strand of RNA and a strand of DNA, in the following manner: Adenines (A) with uracils (U), and the cytosines (C) with the guanines (G). Composition bias—The difference between the sequence o…